Get Complete Project Material File(s) Now! »
MicroRNA-mediated transcriptional and post-transcriptional regulation of CYP genes
MicroRNAs (miRNAs) are a family of small non-coding RNAs of 20-25 nucleotides that have been shown to govern transcriptional and post-transcriptional expression of target genes in many pathways and in a multitude of different organisms (124,125). At present, release 17 of the miRNA database miRBase (http://www.mirbase.org/) shows more than 1000 human miRNAs (126), but the number keeps increasing,; so that it has been estimated that miRNAs are involved in the regulation of expression of 70% of protein-coding genes in human (127). miRNAs are encoded by specific genes. Fifty two per cent of human miRNA-encoding loci are located in intergenic regions and 48% of miRNAs are encoded from intragenic (intronic: 43%; exonic: 5%) regions (128,129). It is, therefore, not surprising that like other parts of the genome, the transcription of miRNA genes is itself under the control of a complex network of several NRs, coregulators and polymerase II (124,128).
In general, miRNAs bind a partially complementary segment within the 3’ untranslated region (3’UTR) of target mRNA to reduce its stability and translation (124). Accordingly, miRNAs could potentially reduce the gene expression, either transcriptionally or post-transcriptionally. Indeed, miRNAs can suppress the transcription of target gene through direct repression of NRs involved in the transcriptional regulation of that gene, leading to an indirect downregulation of target gene transcription. Alternatively, miRNAs can contribute to post-transcriptional gene silencing via direct targeting the 3’UTR transcript of target gene leading eventually to a reduced expression of the target gene. Specifically speaking, miRNAs have been found to act as important transcriptional and post-transcriptional regulators of several CYP genes (130-134). The regulatory effects of miRNA on the expression of CYP genes could be exemplified by the miRNA-mediated transcriptional/post-transcriptional modulation of human CYP epoxygenases. In fact, several NRs that are implicated in the constitutive as well as xenobiotic-induced transcriptional upregulation of CYP2C genes, including PXR, VDR, RXR, HNF4α, ERα and GR, have been found to be regulated post-transcriptionally by various miRNAs (126). The miRNA-mediated post-transcriptional degradation of NR transcripts could be associated with alteration in the NR-mediated transcriptional regulation of CYP2C genes. For instance, miRNA-24 and miRNA-34a negatively regulate the translation of NR HNF4α (135) and, thereby, could indirectly downregulate the HNF4α-mediated transcription of CYP2Cs. Likewise, PXR is regulated by miRNA-148a (136); VDR is regulated by miRNA-125b (137); ERα is regulated by miRNAs 206, 221/222 and 22 (138-140); and GR is regulated by miRNAs-18 and 124a (141). The matter is further complicated by reminding the fact that NRs interact with each other for achieving a precise regulation of CYP2Cs and also have regulatory influence on the transcriptional expression of each other. Additionally, the contribution of miRNAs to the post-transcriptional regulation of human CYP epoxygenases has been the focus of some important recent researches (130-134, 142,143). Notably, CYP2Cs have been recently demonstrated to be downregulated post-transcriptionally by miRNA-103 and miRNA-107, with CYP2C8 being the main target and CYP2C9 and CYP2C19 being affected to a lesser degree than CYP2C8 (142). Likewise, the results of a very recent study revealed that miRNA let-7b reduces CYP2J2 expression so that the decreased expression of this let-7b could lead to the high expression of CYP2J2 protein (143).
NR-epigenetic-miRNA network-mediated regulation of CYP epoxygenases
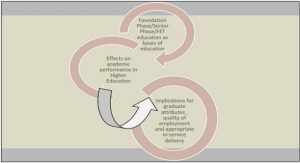
Taking together, in theory, transcription of CYP epoxygenases and consequently EET biosynthesis could be orchestrated by an extremely complex network of NR, miRNAs and epigenetic factors. In this network, CYP expression is primarily induced by NRs, coregulators and Pol II complex. In addition, miRNAs regulate the expression of CYP genes, at post-transcriptional level, by inducing the degradation of CYP epoxygenase mRNAs. Likewise, miRNAs suppress CYP expression, transcriptionally, by repressing the translation of NR/coregulators/Pol II complex. The expression level of all the three main components of this network, i.e. NRs, miRNAs and CYP epoxygenases, is regulated by genetic/epigenetic factors (Figure 10).
Inter- and intra-individual variability in CYP expression
Multiple studies now have demonstrated that there exists a wide inter-individual variability in the expression of CYP mRNA and protein (167-171). There is >50-fold inter-individual variation in the liver enzyme content for CYP2A6, CYP2B6 and CYP2D6; 20-fold for CYP1A2; 12-fold for CYP2E1; and 5-fold for CYP2C and CYP3A4 (167). In addition, highly variable catalytic activities of CYP enzymes have been detected among individuals (167). Likewise, in a given individual, the basal CYP expression level could vary under different physiological and pathological conditions. Herein, we provide a comprehensive overview of the variables that have been identified so far as the potential sources of inter- and intra-individual variability in the CYP expression as well as CYP activity.
Polymorphisms of genes encoding NRs and NR binding sites
The crucial role of NRs and also the importance of NR-RE interaction in the regulation of constitutional and xenobiotic-induced expression of human CYP epoxygenases were discussed thoroughly in section 2.5. Accumulating evidence indicates that mutations in NR binding sites in CYP epoxygenase gene promoters could be associated with dysregulation of these genes (section 2.5). Also, several SNPs have been identified, so far, in genes encoding NRs (106, 231-241). The genetic variations in NR coding regions have been shown to result in altered level of NR proteins (106) and, by this means, can contribute to a variety of local and systemic disorders (242-251), and also to inter-individual differences in drug response through changes in the expression of genes involved in drug metabolism, including CYPs (252-256). For instance, a single nucleotide polymorphism (SNP) in the HNF4-α gene (253), and a genetic variant of the CAR (239), have been identified to be associated with alerted expression of CYP2B6 and several SNPs in the PXR gene may result in CYP3A4 expression and metabolic activity (321,239). Although little is known about the potential influences of NR polymorphisms on the expression and activity of human CYP epoxygenases, considering the importance of NRs in the regulation of these enzymes, it would be logical and makes sense to expect that the genetic variations in genes encoding NRs may potentially be a significant cause of inter-individual variability in the expression of CYP epoxygenases and eventually in the EET formation.
MicroRNAs
As described before, miRNAs regulate the transcription of a wide range of genes such as the xenobiotic-, endobiotic-metabolizing enzymes including CYPs. A growing body of reports has shown that the expressions of miRNAs are readily influenced by xenobiotics, chemicals, carcinogens, drugs, hormones, stress and diseases including cancer, Alzheimer’s disease, CVDs and schizophrenia (126,256-258). This condition-related dysregulation of miRNAs and subsequently the miRNA-related polymorphisms could potentially lead to a considerable inter- and intra-variability in the expression of CYP genes and consequently to changes in the drug and endobiotic metabolism and effects. The miRNA-CYP-mediated variation in drug response would provide useful information in pharmacogenomics and personalized medicine.
Additionally, as discussed, miRNAs themselves are controlled by NR proteins which are, in turn, under control of polymorphic genes (section 3.3.2). The instability in the miRNA expression as the result of NR polymorphisms would provide a potential source of inter-individual variability in the transcription of CYP genes. To complicate matter further, SNPs of genes encoding miRNA precursors, their target sites and silencing machinery were found to alter miRNA function and they are likely to affect phenotypic variation, including disease susceptibility and potentially xenobiotic-, endobiotic metabolism (259,260). In fact, SNPs of miRNA genes may alter their sequences and therefore enhance, diminish or even generate or cancel out their ability to bind to target sites (261-263). Therefore, genetic variations of genes encoding miRNAs could provide an additional source of inter-subject variability in the expression and activity of CYP epoxygenases and in the EET formation.
Epigenetic factors
It has been suggested that there exists a significant inter-individual variability in the epigenetic modification of genes and that the difference in the epigenetic-related regulation of genes could explain a much greater fraction of inter-individual phenotypic variation than differences in genotype, alone (264-268). There is an ever growing body of evidence that environmental factors, including chemical pollutants; dietary components; smoking; temperature changes and other external stressors, can affect the epigenetic landscape of an individual (264-268). Considering the fact that most phenotypic traits are determined by interplay between individual’s gene and environment, the “environmental epigenetics” as a mechanistic link between genes and environment has attracted considerable interest (264). The molecular mechanisms that mediate environment-epigenetic-induced modification of gene expression have been thoroughly reviewed by Feil and Fraga (264).
Additionally, since environmental conditions are dynamic and constantly changing, the environmental epigenetics is being considered a major source of inter- and intra-individual variability in the expression of the genome including genes involved in the metabolism of drugs and endogenous compounds such as CYPs (269-273). The study of epigenetic-mediated inter- and inter-personal differences in the expression of drug-metabolizing enzymes is a rapidly emerging discipline called “pharmacoepigenetics”; it is expected that pharmacoepigenetics, complementary to pharmacogenetics, will play a crucial role in future pharmacology and personalized medicine (269-273).
Similarly, the regulatory impact of epigenetic variations and the environmental epigenetics on human CYP epoxygenase genes, specifically CYP2C19 and CYP2J2 (section 2.7), may result in a significant inter- and intra- variability in the expression of epoxygenases and eventually in the biosynthesis and biological function of EETs.
CYP gene polymorphism
The human CYP genes are highly polymorphic (7,34,40,155,274). The polymorphisms can modify the function of CYPs through several mechanisms: loss-of-function polymorphisms mostly affect splicing and expression, but surprisingly not transcription or protein structure (7,275); gain-of-function polymorphisms influence enzyme expression and activity via CNV, promoter variation (e.g. CYP2B6 and CYP2C19) (7), and amino acid variations that resulting in increased substance turnover (e.g. CYP2B6 and CYP2C8) (276). Unexpectedly, from all the discovered polymorphisms, there are few that influence on substrate selectivity and xenobiotic-mediated inducibility of CYP enzymes (7).
CYPs represent the most important phase I drug metabolizing enzymes and genetic polymorphisms within CYPs remarkably affect the metabolism of drugs that are substrates for those particular enzymes which eventually leading to differences in drug response and risk of adverse drug reactions. Very recently, the characteristics, functional effects and clinical implications of pharmacogenetically important CYP polymorphisms have been comprehensively by Zanger and Schwab (7).
Additionally, since EETs, as epoxygenase-mediated AA metabolites, possess potent protective effects on the cardiovascular system, genetic variations in the CYP epoxygenase pathway have attracted considerable interest.The main human CYP epoxygenases including CYP2C8, CYP2C9, CYP2C19 and CYP2J2 exhibit genetic polymorphisms. The most common epoxygenase gene polymorphisms (Table 1) have been demonstrated to be associated with a significant decrease in their epoxidation activity and consequently in the EET biosynthesis (34). For instance, CYP2C8*3 has shown to be defective in the metabolism of AA to 11,12-EET and 14,15-EET having a turnover of 35-40% of wild type CYP2C8 (CYP2C8*1) (34). Likewise, CYP2C9*2 displays half of CYP2C9 (CYP2C9*1) wild type epoxygenase activity (34). Moreover, compared to other CYP2C8/CYP2C9 haplotypes, samples genotyped asCYP2C8*3/*3/CYP2C9*2/*2 exhibited 34% decreased EET biosynthesis (34). Similarly, CYP2J2*7 (G-50T) polymorphism, the most frequent and the most studied functional variant of CYP2J2, results in a reduced transcription of the CYP2J2 gene and also lower EETs plasma concentration (34). Considering the fact that CYP epoxygenase polymorphisms cause lower activity of these enzymes and that EETs play important roles in cardiovascular physiology, it is logical to assume that genetic variations in the epoxygenase pathway could be generally associated with an increased risk of several CVDs including HTN and CAD and, thereby, could be considered one of the determinants of individual susceptibility to these disorders (34,40).
As demonstrated in Table 1, the frequencies of CYP epoxygenases polymorphisms are highly dependent on the ethnic background (7,34). While the frequency of CYP2C8*2 has been reported to be up to 18% in African-Americans (rare in Caucasians), CYP2C8*3 is primarily occurred in Caucasians (up to 19.8%) and neither is occurred in Asians. Furthermore, CYP2C9*2 and CYP2C9*3 are considerably frequent in Caucasians (up to 19% and 16.2%, respectively), while these polymorphisms are less frequent in African and are rare in African-American and east Asians. Likewise, the frequency of CYP2C19*2 has been reported to be 45.5%, 7.8% and 13% in Chinese, Bolivian and Caucasians, respectively. There is also a large inter-ethnic difference in the prevalence of CYP2J2*7 polymorphism, seemingly being most frequent in Africans (7,34).
CYP expression in cardiocirculatory system
The tissues in the cardiovascular system are the main target organs of numerous widely-prescribed medications such as β-blockers, angiotensin converting enzyme inhibitors, angiotensin receptor blockers, calcium channel blockers and anticoagulant agents; the metabolic activity of cardiocirculatory CYPs involved in drug metabolism may have a significant effect on the effectiveness of pharmacotherapy (412-415). For example, the expression of CYPs in the right ventricle has been found to influence the metabolisms of β-blockers and verapamil and may explain, at least in part, lack of efficacy of these drugs in certain patients (412-414). Moreover, the cardiac CYPs have a potential role in the detoxification of noxious foreign compounds (412-414). Additionally, strong evidence indicates that CYPs expressed in the cardiocirculatory tissues also contribute to the biotransformation of endogenous compounds such as AA (i.e. to EETs) (45,56-60,390,414-417). Considering the facts that EETs act, predominantly, in an autocrine and paracrine manner, and that EETs have potent cardiovascular protective properties, the cardiovascular CYPs may have a clinically important impact on the synthesis and biological activities of endobiotics, particularly EETs.
Table of contents :
CHAPTER 1 – INTRODUCTION
I. Pharmacogenomics of Anti-inflammatory Effects of Thienopyridines
1. Pharmaco-genetics, -genomics and personalized medicine
2. P2Y12 receptor antagonists including thienopyridines
2.1. Clopidogrel
2.1.1. Anti-inflammatory effects of clopidogrel
2.2. Prasugrel
2.2.1. Anti-inflammatory effects of prasugrel
3. References
II. Influences of Inter-tissue, Inter-individual and Intra-individual Variability in the Expression of Human Cytochrome P450 Epoxygenases on Epoxyeicosatrienoic Acids-m
1. Introduction
2. Cytochrome P450, Epoxyeicosatrienoic acids; General considerations
2.1. CYP enzyme; structure
2.2. CYP enzyme; catalytic activity
2.3. Human CYP epoxygenases; EET formation
2.4. Physiological roles of EETs in vascular biology
2.5. Nuclear receptor-mediated transcriptional regulation of CYP epoxygenase genes
2.6. MicroRNA-mediated transcriptional and post-transcriptional regulation of CYP genes
2.7. Epigenetic-mediated transcriptional and post-transcriptional regulation of CYP genes
2.8. NR-epigenetic-miRNA network-mediated regulation of CYP epoxygenases
2.9. Genes counter-interaction
3. Inter- and intra-individual variability in CYP expression
3.1. Age
3.2. Gender
3.3. Molecular genetics
3.3.1. Variation in POR expression
3.3.2. Polymorphisms of genes encoding NRs and NR binding sites
3.3.3. MicroRNAs
3.3.4. Epigenetic factors
3.3.5. CYP gene polymorphism
3.3.5.1. Copy number variation
3.4. Xenobiotics
3.5. Environmental factors
3.6. Sex steroid-dependent states
3.7. Immunological response – local/systemic disorders
3.8. Nervous system and stress mediated CYP regulation
3.9. Radiation
4. Inter-tissue variability in CYP epoxygenase expression
4.1. CYP expression in liver
4.2. CYP expression in gastrointestinal tract
4.3. CYP expression in respiratory tract
4.4. CYP expression in genitourinary system
4.5. CYP expression in central nervous system
4.6. CYP expression in cardiocirculatory system
5. Inter-tissue, inter- and intra-individual variability in CYP epoxygenases; clinical implication
6. Summary
7. References
III. Influence of Inflammation on Cardiovascular Protective Effects of Cytochrome P450 Epoxygenase-Derived Epoxyeicosatrienoic Acids
1. Vascular arachidonic acid metabolism
2. EETs synthesis and metabolism; overview
3. CYP epoxygenases expression and EET resources in cardiovascular system
4. Physiological roles of EETs in cardiovascular system
4.1. EETs and myocardial function
4.2. EETs and vascular function
4.2.1. EETs and vasodilation
4.2.2. EETs and vascular homeostasis
4.2.2.1. EETs and regulation of angiogenesis, fibrinolysis, apoptosis and SMCs migration
4.2.2.2. CYP epoxygenase-derived EETs and inflammation
4.2.2.2.1. CYP epoxygenase-derived EETs and inhibition of EC activation
4.2.2.2.1.1. NF-κB and inflammation
4.2.2.2.1.1.1. NF-κB; structure and activation pathways
4.2.2.2.1.1.2. CYP epoxygenase-derived EETs as NF-κB inhibitors
4.2.2.2.1.2. CYP epoxygenase-derived EETs and activation of PPAR and EGF receptor
4.2.2.2.1.3. CYP epoxygenase-derived EETs and inhibition of LOX-5 and COX-2
4.2.2.2.1.4. CYP epoxygenase-derived EETs and increase HO-1 expression
4.2.2.2.1.5. CYP epoxygenase-derived EETs and increase eNOS expression .
4.2.2.2.2. CYP epoxygenase-derived EETs and platelet inhibition
4.2.2.3. Effect of inflammation on CYP epoxygenase expression
5. Inflammation, CYP epoxygenase expression and EET generation; a vicious cycle?
6. Conclusion and perspective
7. References
IV. Point-of-care pharmacogenetic testing
CHAPTER 2 – HYPOTHESES AND OBJECTIVES
CHAPTER 3 – POPULATIONS AND METHODS
1. SC
1.1. Sample reference population
1.2. Genotyping
1.3. Hp measurement
2. Populations on Thienopyridines
2.1. Paris Study
2.1.1. Measurement of inflammatory markers
2.1.2. Genotyping
2.2. Marseille Study
2.2.1. Biochemical profile
2.2.2. Genotyping
2.2.3. Platelet function measurement
2.3. Liverpool Study
CHAPTER 4 – RESULTS
References
Association between CRP Levels and High on-treatment Platelet Reactivity in Post-PCI on-Thienopyridine Patients
CHAPTER 5 – GENERAL DISCUSSION AND PERSPECTIVES
I. Thesis main findings
II. Inflammation, CVD and post-PCI ISR
1. Platelets: key players in vascular inflammation
1.1. Platelet interaction with ECs
1.1. Platelet interaction with leukocytes
1.2. Activated platelets and CRP, Hp and orosomucoid acid
2. Inflammation and post-PCI complications
2.1. ISR after DES vs. BMS
III. CYP and Metabolism of EETs and Thienopyridines
IV. Thienopyridine Resistance, Non-responsiveness and High On-treatment Platelet Reactivity
1. Resistance, hypo-, non-responsiveness to thienopyridines
2. High on-treatment platelet reactivity
3. Resistance vs. high on-treatment platelet reactivity
V. Smoker’s paradox
VI. Necessity of experimental studies?
VII. Perspectives
2.1. EET measurement:
VIII. References
CHAPTER 6 – Résumé de Thèse En Français
A. Introduction
B. Hypothèses et objectifs
C. Populations
D. Résultats
E. Discussions
F. Perspectives