Get Complete Project Material File(s) Now! »
Cell metabolism and Phospholipid biosynthesis
Outline
This Chapter is mainly focused on the cell metabolism as a vital cellular process and on the phospholipids which are the major component of biolog-ical membranes. We next present a graph implementation for the core of Glycerophospholipid metabolism in which the pathways are supplied from bibliographical references and some online databases. [15, 16, 17, 18, 19, 20]
Structure and processes in a cell
The cell surrounded by a lipid membrane is the smallest functional basic unit of life. A cell contains all the components required for its replication and this is the reason to be distinguished from smaller biological units. There are two types of cells: eukaryotic (with nucleus) and prokaryotic (without nucleus). Prokaryotic cells are relatively small in size and independent, while eukaryotic cells, which are typically larger than prokaryotic cells, are usually found in multicellular organisms. Diverse biological processes such as growth, metabolism and replication are carried out by prokaryotic and eukaryotic. All cells, whether prokaryotic or eukaryotic, have a membrane that envelops the cell, separates its interior from its environment, regulates what moves in and out, and maintains the electric potential of the cell.
Metabolism
Metabolism is a vital cellular process that consists of the set of chemical reactions that happen in living organisms to maintain life. Since the mal-function in metabolism is a major reason of human disease, it is important to construct and study metabolic networks. That is to say, the sets of co-herent processes that organize metabolic networks are complex and highly interconnected, therefore there is a need to the computational approaches (see Chapter 1 ). The reconstruction of the metabolic networks and kinetic modeling is the core of systems biology. [21, 22]. One can use the metabolic network to suggest potential alternatives to drug targets or study the e ects and causes of diseases like cancer and copnsequently hints for tentative ther-apies. Next we formulate the metabolic network into a mathematical model based on rate laws for enzymatic or non enzymatic reactions. This kinetic model can become quite complex with a large quantity of parameters.
Metabolism is de ned as the totality of the chemical reactions catalyzed by enzymes that are carried out in an organism. The changes in metabo-lites are called biotransformations and a metabolic pathway is a sequence of biotransformations [23] as illustrated in gure 2.1.
Figure 2.1. Multi-enzyme reaction: metabolic pathway where multiple biotransformations take place. The substrate of each reaction is the product of the previous reaction.(See [23])
In a metabolic pathway the input metabolites are called substrates and output metabolites are called products. Metabolites can originate from food, but can also be products of other metabolic pathways in the cell. The phenotype of the cell is determined by its metabolism. One can improve the cellular properties by changing the regulation of metabolic pathways through changes in enzyme and metabolite concentrations. Enzymes, metabolites, their respective interactions and the reactions involved in these pathways have been studied by many researchers and are stored in di erent databases, e.g.:
Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway data-base. (http://www.genome.ad.jp.kegg/pathway.html)
BRENDA. (http://www.brenda-enzymes.info/) a Biochemical Genetic and Genomic knowledgebase of large scale metabolic reconstruction (BiGG). (http://www.bigg.ucsd.edu)
KEGG is a collection of online databases dealing with genomes, enzy-matic pathways and biological chemicals. The pathway database of KEGG records networks of molecular interactions in the cells and variants of them speci c to particular organisms. The KEGG database can be used for mod-eling and simulation, browsing and retrieval of data. Such data bases are essential in the systems biology approach.
Phospholipids
Phospholipids are a major component of biological membranes.(Fig. 2.2) They are a class of lipids formed from four components: fatty acids, a negatively-charged phosphate group, alcoholamine and a backbone. Phos-phatidylCholine (PtdCho) and PhosphatidylEthanolamine (PtdEth) are two of the most abundant phospholipids. Biosynthesis and metabolism of these and other phospholipids are important for proliferation of membrane-bound organelles, lipoprotein synthesis, and signal transduction a ecting processes of cell proliferation, di erentiation and apoptosis. Yet our understanding of this metabolism and its regulation is far from complete.
It has long been recognized that di erent cell types and tissus display unique and stable pro les of PtdCho and other phospholipid species. Perturbation of PtdCho homeostasis in mamalian cells leads to cell death. In the early 1970s Sundler et al. [24, 25] examined the rates of synthesis for liver PtdCho and PtdEth using radioisotope methods. Their data could not be described by a simple precursor-product relationship and raise questions about the compartment of metabolite pools and/or channeling of metabolic pathways which are yet to be fully answered. Additionally, many questions about the metabolic pathways themselves remain unanswered.
In most eukaryotic cells, PtdCho is synthesized through two di erent path-ways [26]; in the cytidine diphosphate-choline (CDP-choline) pathway (also known as the Kennedy pathway) and via the transmethylation of PtdEth catalysed by PE-N-methyltransferase (PEMT). Choline, supplied by food, is principally in the form of PtdCho but also exists as free Choline [27, 28]. Choline is an essential nutrient for all cells because it plays a role in the syn-thesis of the phospholipid components of the cell membranes, as a methyl-group donor in methionine metabolism. Quantitatively, PtdCho is the most important metabolite of Choline and accounts for approximately one half of the total membrane lipid content.
Figure 2.2. Cell membrane structure: proteins and phospholipids are major components of biological membranes. (Image source: http://commons.wikimedia.org/wiki/File:Cell membrane detailed diagram
The Kennedy pathway for producing PtdCho, involves the activation of Choline (Cho) to CDP-choline through an intermediate product, Phospho-Choline (P-Cho). The second pathway to produce PtdCho consists of three sequential methylations of phosphatidylethanolamine. Cho derived from the turnover of PtdCho produced by the methylation pathway is used for Ptd-Cho synthesis through the Kennedy pathway. Therefore the activity of the Kennedy pathway does not reduce even in the absence of Cho in the growth medium [29].
In 1975, Sundler et al. used radioisotope methods to examine the rates of synthesis for PtdCho and PtdEt of liver [30, 31]. In their study, there are still questions about compartmentation of metabolite pools and channeling of metabolic pathways to be answered. However evidence of the two di erent pathways of PtdCho synthesis and the relative activities of these pathways was provided by Vance et al. [32, 33]. The Nuclear Magnetic Resonance (NMR) spectroscopy method has been used to study the biosynthesis of PtdCho and PtdEth [34, 35]. The NMR technique can also provide a de-tailed examination of the speci c metabolic pathways. Reo et al. performed kinetic analyses of liver PtdCh and PtdEth biosynthesis using 13C NMR spectroscopy [36].
The development of methods for pathway-speci c analyses of phospholipid biosynthesis in intact tissue can help in our understanding of numerous cel-lular processes, and may be important for cancer studies. This is why the Phospholipid metabolism has attracted the attention in cancer research. It is of interest to biologists to be able to follow the phospholipid metabolism in circumstances in which cell survival and cell proliferation are of concern, e.g. neurological disorders and cancer [37, 38]. Thus there is a need to develop a model for their biosynthesis and turnover. This is why we tried to nd a model for the core of GlyceroPhospholilid metabolism. Our goal is to build a model with which one could simulate the behavior of phospholipid interactions taking into account the impact of the environment. Due to the complexity of this system, mathematical modeling and numerical simulation are necessary to enable a compact representation of the current knowledge and to make meaningful quantitative predictions guiding future experimen-tal studies.(See section 3).
Biochemistry of the phospholipid metabolism
Figure 2.3 illustrates the model of Glycerophospholipid metabolism. The model of phospholipid metabolism that we describe here is the core of Glyc-erophospholipid metabolism which is supplied from bibliographical refer-ences(e.g. M.Israel and L.Schwartz [15, 16, 17, 18, 19, 20]).
Our analysis concerns twenty-four biochemical reactions, as illustrated in Fig.2.4. In this system there are two main sub-systems, which have al-most the same reaction structures; the rst one is the Choline (Cho) cycle and the second one is the Ethanolamine (Eth) cycle. In order to have a more complete model several reactions involving external reactants are also considered and studied in the model; In gure 2.4 blue and green arrows rep-resent Choline cycle and Ethanolamine cycle respectively and Pink arrows correspond to reactions which connect these two sub-cycles. The chemical structures of metabolites are shown in gure 2.5.
2.4.1. Choline cycle. Cho is phosphorylated in an enzymatic reaction catalyzed by Choline-Kinase (CK), resulting in the formation of Phospho-Choline (PC)[16]. PC is converted to PhosphatidyleCholine (PtdCho) in a two step reaction, rst catalyzed by regulatory enzyme PhosphoCholine-Cytidyl-transferase (CCT), then by PC-transferase (CTP)[16, 17]. PtdCh is converted to Glycero-PhosphoCholine (GPC) in the reaction catalyzed by Phospholipase A2 (PlpA2)[17]. In addition, PC and Cho can be synthesized from hydrolysis of PtdCho through the reactions catalyzed by Phospholipase
Arrows with V Mi and KMi parameters refer to enzymatic reactions while the others represent simple reactions. Blue and green arrows represent Choline cycle and Ethanolamine cycle repectively and Pink arrows correspond to reactions which connect these two sub-cycles. Reactants: Cho (Choline), PC (Phospho-Choline), PtdCho (Phosphatidyle-Choline), GPC (Glycero-PhosphoCholine), Eth (Ethanolamine), PE (Phospho-Ethanolamine), PtdEth (Phosphatidyle-Ethanolamine), GPE (Glycero-PhosphoEthanolamine). En-zymes: CK (Choline-Kinase), EK (Ethanolamine-Kinase), CCT/CPT (PhosphoCholine-Cytidyl-Transferase), EC-T/EPT (PhosphoEthanolamine-Cytidyl-Transferase), PEMT (PhosphatidyleEthanolamine-N-methyl-Transferase), PlpA2 (PhosphoLipase A2), PlpC (PhosphoLipase C), PlpD (PhosphoLi-pase D). Parameters: VM (Michaelis maximum reaction rate), KM (Michaelis concentration constant), ki(Rate constants for external reactions).
C (PlpC) and Phospholipase D(PlpD) respectively[17, 18]. Cho can be also synthesized from GPC[18].
Ethanolamine cycle
Eth is phosphorylated in an enzymatic reaction catalyzed by Ethanolamine-Kinase (EK), resulting in the forma-tion of PhosphoEthanolamine (PE)[19]. PE is converted to Phosphatidyle-Ethanolamine (PtdEth) in a two step reaction, rst catalyzed by the regula-tory enzyme PhosphoEthanolamine-Cytidyl-transferase (ECT), then by PE-transferase (EPT)[19, 20]. PtdEth is converted to Glycero-PhosphoEthanolamine (GPE) in the reaction catalyzed by Phospholipase A2 (PlpA2)[33, 17]. Eth is synthesized from GPE[18].
The above two sub-systems are related through the reaction between Pt-dEth and PtdCho where PhosphatidylEthanolamine N-Methyl Transferase (PEMT) plays the role of catalyst[29, 32, 33]. This reaction seems to be an important reaction in this system, and is the basis of the main analysis in our study, since the homeostasis of PtdCho is essential to maintain cell survival.
External reactions
In addition to the reactions described so far, most of the reactants in phospholipid metabolism models have external reactions. For example there is a reversible reaction in which Phosphatidyle-Serine (PtdSer) releases CO2 and PtdEth as products [39]. In the same way there are several external reactions in which Cho, Eth, PC, PE, PtdCho and PtdEth have the role of substrate or product. We present these external re-actions by input or output arrows in the model(Fig.2.4)[34, 39, 16].
Table of contents :
Acknowledgements
Abstract
Résumé
Chapter 1. Introduction
1.1. Computational biology
1.2. Systems biology
1.3. Modeling and Simulation
1.4. Kinetic modeling
1.5. Computational systems biology in cancer
1.6. The studied biological model
1.7. Manuscript Plan
1.8. Main results
Chapter 2. Cell metabolism and Phospholipid biosynthesis
Outline
2.1. Structure and processes in a cell
2.2. Metabolism
2.3. Phospholipids
2.4. Biochemistry of the phospholipid metabolism
Chapter 3. Mathematical modeling of phospholipid biosynthesis
Outline
3.1. An introduction to the mathematical modeling of metabolic networks
3.2. The mathematical translation of metabolic networks
3.3. Modeling process
Chapter 4. Parameter estimation, Simulation and Stability analysis
Outline
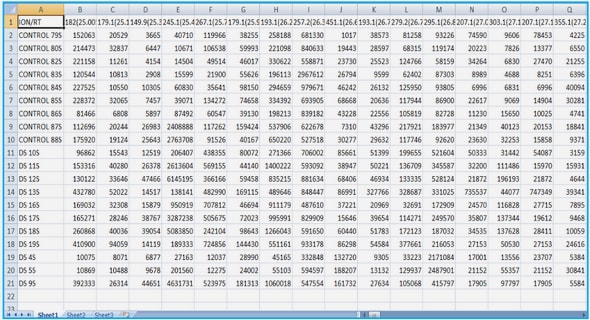
4.1. Where to get data from?
4.2. Parameter estimation
4.3. Simulation
4.4. Stability analysis
Chapter 5. Model’s application to healthy rat’s liver
Outline
5.1. Concentrations and Parameter estimation
5.2. Phase spaces
5.3. Stability analyses
5.4. Stability analyses results
5.5. Complexity study
5.6. Mathematical proof of stability (sketch)
5.7. Conclusion
Chapter 6. Model’s application to B16 melanoma and 3LL carcinom cells in response to CENU
Outline
6.1. Model application and Parameter estimation
6.2. Comparative analyses of parameters
6.3. Results
6.4. Conclusion
Chapter 7. Model’s application to B16 melanoma cells in response to methionine deprivation
Outline
7.1. Mathematical model and parameter estimation
7.2. Comparative analyses of parameters
7.3. Sensitivity analysis
7.4. Conclusion and Discussion
Chapter 8. Conclusion
Outline
8.1. Summary and main results
8.2. Discussion and Future work
Appendix A. Chemical reactions and enzyme kinetics
Outline
A.1. Chemical Reactions
A.2. Reaction rate
A.3. System of chemical reactions
A.4. Enzyme kinetics
Appendix B. Parameter estimation methods
B.1. Forward or bottom-up modeling
B.2. Using steady-state data
B.3. Inverse or top-down modeling
Appendix C. Stability analysis in dynamic models
Outline
C.1. Stability Analysis of Dynamic Models
Appendix D. Experimental data of CENU
D.1. B16 melanoma and 3LL carcinoma cells in response to CENU
Appendix E. Experimental data of MDS
E.1. B16 melanoma cells and response to methionine deprivation (MDS)
E.2. Biological global eect of MDS
Appendix F. Basis for a new software for biological networks modeling
Outline
F.1. Introduction
F.2. Goal
F.3. Data and Methods
F.4. Development tools
F.5. Graph construction
F.6. Analyses
F.7. Rate laws to construct the equations
F.8. Inhibitors
Bibliography