Get Complete Project Material File(s) Now! »
The process of epidemic intelligence: from data collection to reporting
To detect health threats in a timely manner and monitor their evolution, EI activities are based on the systematic review of a variety of information sources, both official and unofficial.
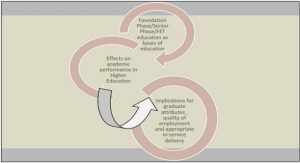
The EI procedure is divided into several steps: data collection (including screening and filtering), verification, analysis and communication to health authorities and decision-makers (Figure 3). These steps are usually formalized through an SOP (standard operating procedure). Depending on the situation, these steps can take time, particularly the validation and interpretation of the signal as they might involve several experts who might not be available to react instantly. Despite the number of steps in the EI procedure and the many people it might involve, the goal is to validate all the steps from data collection to reporting rapidly in order to ensure timely detection and communication of health threats to authorities and, in turn, timely action.
A signal is defined as any health-related information collected by the EI system, which could potentially impact human or animal populations (depending on the scope of the system). An event is a signal that has been selected (i.e. represents a potential threat according to criteria defined by health authorities) and verified.
Different definitions exist for a signal and an event. I choose to use the terms as currently defined by ECDC and WHO, and not the definitions from Paquet et al. which are inversed, i.e. a signal corresponds to a verified event.
If the event is analyzed and judged to require the implementation of control or prevention measures by health authorities, it becomes an alert.
Data collection
When setting up EI activities, it is important to define the mandate and the scope of the system by determining the risk group. In the case of the French epidemic intelligence system (FEIS) which I led for four years, the EI team monitors any health-related event that could have a potential impact on the animal population in France, with a main focus on livestock (poultry, cattle) but also including wild animals, pets and other domestic animals.
Health-related information can be collected from many different information sources, both official and unofficial. Considering multiple sources of information, particularly combining unofficial and official sources, has been shown to improve the timeliness of the detection and the accuracy of the information collected (Yan et al., 2017a). For animal health, the main official sources of information for international EI activities are the Empres-I database from the Food and Agriculture Organization (FAO), the World Animal Health Information System (WAHIS) of the OIE and the Animal Disease Notification System (ADNS) of the European Commission. For human health, some examples of official information sources are the WHO, the ECDC and the Center for Disease Prevention and Control (CDC). Official information sources also include national or sub-national governmental health agencies, whether for veterinary or human health. Examples of unofficial sources for both veterinary and human health include media articles, rumors and unstructured personal communications from experts. The scope of sources monitored remains flexible and can rapidly and easily be adapted to the epidemiological context, for instance by strengthening EBS and monitoring local media outlets more closely for a country where a suspected outbreak has been detected. Once the data has been screened, it is reviewed and filtered in order to detect new or updated information concerning potential health-related threats. Relevant information regarding health events are for example the occurrence of an outbreak of an EID, a significant increase in the endemic circulation of a known infectious disease, reports of high mortality or morbidity in animals with an unknown etiological cause, or epidemiological updates on ongoing EID outbreaks. Data filtering allows experts to discard information out of the surveillance scope (e.g. benign or non-infectious diseases), false rumors and duplicates in the collected data. This step is closely linked to the mandate of the institution and the scope of the EI system.
The French animal health epidemiological surveillance platform
In France, the main challenges in animal health have evolved in the past decades with recent emergences leading to significant economic losses. Examples include bluetongue serotype 8 in 2006 (Sailleau et al., 2017), Schmallenberg virus in 2011 (Dominguez et al., 2014), and highly pathogenic avian influenza H5N8 in 2016 (Guinat et al., 2018). Thankfully, surveillance, prevention and control efforts have improved in the past years. Following the acquisition of a disease-free status for multiple contagious diseases (e.g. FMD or rabies), the focus of disease surveillance in France switched to vigilance. Instead of monitoring the evolution of diseases already present in the country, health authorities had to focus on the early detection of new and emerging diseases at the international level to anticipate new introductions of emerging health threats.
Existing surveillance systems needed to be adapted in order to face the new challenges of disease emergence. For example, setting up a non-disease-specific surveillance through the implementation of syndromic surveillance would cover a broader scope of diseases with similar clinical manifestations. Partly due to these considerations, the French Ministry of Agriculture suggested the creation of a national platform for epidemiological surveillance in animal health. The French animal health epidemiological surveillance (ESA) platform1 was set up in 2011 to improve the efficiency of epidemiological surveillance at national level (Figure 4).
The ESA Platform counts six founding members, with four additional members added by 2018. These members represent different sectors of animal health: the General Directorate for Food (DGAL, part of the Ministry of Agriculture), farmers (La Coopération agricole, GDS France), veterinarians (French society for veterinarian technical groups – SNGTV), scientific support organizations (the Agency for Food, Environmental and Occupational Health & Safety – ANSES, CIRAD and the National Research Institute for Agriculture, Food and Environment – INRAe), laboratories (French association of public veterinary laboratories – Adilva), hunters (National Hunters’ Federation – FNC), and wildlife and biodiversity services (French Office for Biodiversity – OFB) (Figure 5).
Through its members, the ESA Platform provides scientific support to improve animal health surveillance efficiency through a public/private partnership.
Epidemic intelligence activities
The FEIS combines IBS and EBS by monitoring both official (e.g. French animal health authorities, OIE, FAO, the European Commission) and unofficial (e.g. media, ProMED-mail, personal communication) sources of information relating to animal health. Once collected, the information is verified and analyzed with the FEIS’ network of national and international experts (Figure 8). This allows the FEIS to analyze and interpret the event according to the context when writing a report, and include information relating to the viral strain, seasonality patterns, the previous occurrence or the historical circulation of the virus in the country. Information relating to different disciplines, such as epidemiology, virology or entomology, is also useful to provide context and a holistic view taking into account complementary perspectives. This reinforces the need for a pluridisciplinary approach to EI, which is why the FEIS team and expert network include experts from a wide range of disciplines such as epidemiology, informatics, entomology, virology, and many other specialties.
Preliminary evaluation of the FEIS
The contribution of the FEIS to the EI process (data collection, verification, analysis and reporting) have been presented in a scientific article. After three years of leading and improving the FEIS, I decided to conduct a short study to evaluate if the FEIS detected and reported on all important events relating to animal health according to its scope, as a way to test its efficiency. To do so, I compared the reports produced by the FEIS to posts published by ProMED from 1st January 2016 to 31st December 2017, to identify if the system had missed any information. I set up a partnership with ProMED directors (based in the United States of America) and moderators in order to promote international collaborations through the bilateral exchange of information and discussions relating to methodology. The results of the analysis of the reports published by the FEIS from 2016 to 2017, and the comparison of these reports to ProMED alerts are presented in Mercier et al., 2020.
Limitations and challenges
EI systems face several technical and organizational challenges that can hamper the exhaustiveness, timeliness and efficiency of these systems:
Completeness of the collected information and its ability to reflect the current epidemiological situation is a main challenge. As mentioned, traditional surveillance systems relying on IBS to gather validated and official information in the form of indicators. However, this method may miss information in the early stages of disease emergence, and lacks timeliness and sensitivity in the detection of unusual events relating to emerging diseases. To overcome this challenge, some EI systems have set up an EBS component and use automated biosurveillance tools to monitor unofficial information sources such as online media reports or scientific publications.
Timely verification of the information collected, particularly if the information originates from an unofficial source like the media or rumors, is another challenge. As previously stated, unofficial information can be validated using official sources or a network of experts. In the framework of the FEIS, this challenge was addressed by setting up an extensive network of national and international experts with different fields of expertise (e.g. epidemiology relating to both human and animal health, virology, entomology, farming systems). This network includes all of the French national reference laboratories for animal diseases as well as several European and OIE reference laboratories, which can be a very useful source of validation of information regarding the occurrence of an emerging disease.
Timely reporting of health threats to authorities in charge of response can also be challenging. The internal hierarchical chain of validation of reports can delay the dissemination of information, particularly in the case of large institutions or agencies. To address this issue, French animal health authorities have set up the ESA Platform to gather animal health professionals and stakeholders from different sectors (veterinary agencies, farmers, veterinarians, wildlife services, hunters, laboratories and research centers) and facilitate the exchange of information, methods and expertise.
International EI activities conducted by international organizations such as WHO, OIE or ECDC have their limits, but can be complemented by national-level EI systems. International systems provide a global coverage and monitor a large variety of information sources, which makes them very efficient at timely detection of health risks. The analysis of the threat, however, is very country-dependent. This is why some countries have set up an international EI system that uses international systems as information sources, but also provide an interpretation and analysis tailored to the country’s needs. This means that health threats are filtered depending on the level of risk they represent for the specific country. This risk depends on several factors such as historical disease circulation, economic ties (e.g. trade, governance), travel access or geographical proximity. This challenge was addressed in France with the creation of a French international EI system as part of the French animal health epidemiological surveillance platform.
The integration of new methods, data sources or tools can be complicated by several factors. First, we should aim to improve and complement the existing systems already in place instead of creating new and independent tools, which can duplicate and complicate the work. Also, the developed methods or tools should fit the needs of end-users in order to ensure appropriation and sustained use of the tool. We are addressing these challenges in the framework of the H2020 MOOD project proposal by integrating a thorough review of existing activities, systems and tools, and adding a social science component in one of the work packages. This component aims to identify the main gaps and challenges relating to EI data and work flows within institutions of EI activities, and identify end users’ needs in terms of data, tools and methods for the co-conception of solutions. The integration of social science processes such as participatory methods can facilitate interactions and communication in research projects involving many different stakeholders, disciplines and geographical scales.
Platform for automated extraction of animal disease information from the web (PADI-web)
Although many biosurveillance tools exist and have proven their efficiency in detecting outbreak news on the web, they provide limited coverage of animal diseases (Valentin et al., 2020b). The Platform for Automated extraction of animal Disease Information from the web (PADI-web) is an automated text mining platform that detects, categorizes and extracts disease outbreak information from news articles published on the Internet (Figure 10) (Arsevska et al., 2018). PADI-web was designed for the FEIS to facilitate the monitoring of animal disease outbreaks from unofficial sources of information such as online news articles. It incorporates intelligent systems based on text mining which include natural language processing, machine learning and data mining techniques (Arsevska et al., 2018).
A first module of the platform aims to identify and collect online news articles using a series of customized Really Simple Syndication (RSS) feeds on Google News. Google News was chosen as the main data source because it is freely accessible and allows users to tailor search parameters. The RSS feeds use a list of predefined keywords relating to hosts, symptoms and disease names (Arsevska et al., 2016). They can be disease-specific (using disease names) or non-specific (using a combination of symptoms and hosts). This allows the tool to monitor nine specific diseases of interest (ASF, classical swine fever, avian influenza, FMD, bluetongue, Schmallenberg virus, lumpy skin disease, peste des petits ruminants and West Nile), as well as information on other diseases or syndromes (e.g. early stages of emergence, before official confirmation of etiology).
The majority of RSS feeds use English keywords, but we have also implemented feeds in other languages to increase local media coverage. These languages were selected to target areas with enzootic circulation of specific diseases or at high risk of disease emergence (e.g. adding Arabic to monitor FMD in endemic Arabic countries, or Chinese to monitor ASF). The integration of multilingual RSS feeds has significantly increased the number of relevant news articles detected (Valentin et al., 2020b).
Once collected, news articles are processed to remove duplicates from the database. News articles that were retrieved using non-English RSS feeds are translated into English. News items are then classified to determine if they are pertinent, i.e. relating to an animal disease event (e.g. describing a current outbreak, prevention and control measures, or socioeconomic impact of a disease). This classification step is run by a supervised machine learning process. In this process, experts define two classification categories (pertinent/non pertinent) and manually annotate a small dataset in order to build a model (supervised classification). Based on labeled data, the machine-learning approach takes into account textual content of both classes in order to construct the classification model. This model is then able to establish definitions for the two classes based on the expert-annotated dataset, and annotate new data (machine learning). The model is continuously enriched as new data is added to the database and classified (daily update of the model). This underlines the importance of expert annotation and input at the beginning of the process to set the foundation for the model classification, and to continuously update the tool’s keywords if new disease threats emerge. A more specific type of classification has been recently implemented, and consists of five topic categories to go beyond binary relevance classification (Valentin et al., 2020c):
confirmed outbreak: information about a new or ongoing confirmed outbreak.
suspected outbreak: information about new or ongoing cases which have not yet been diagnosed but are associated with a suspected disease.
unknown outbreak: information about new or current cases not yet diagnosed and not associated with a suspected disease.
preparedness: information about prevention measures in a country not yet affected by the diseases but on alert to prevent the introduction of the disease, and.
impact: information on economic, societal or political impact of disease outbreaks in an affected country or region.
Table of contents :
Introduction
1.1. Objectives
1.2 Context
Epidemic intelligence
2.1. Definition
2.2. Elements of language
2.2.1 Risk and hazard
2.2.2 Indicator and event-based surveillances
2.2.3 The process of epidemic intelligence: from data collection to reporting
2.2.4 Existing systems
2.2.5 Presentation of the French epidemic intelligence system for animal health
2.3. Preliminary evaluation of the FEIS
2.4. Limitations and challenges
A pluridisciplinary approach to data collection in the context of tool development
3.1. Introduction
3.2. Development of an automated media monitoring tool for animal health (PADI-web) .
3.2.1 Internet event-based surveillance
3.2.2 Platform for Automated extraction of animal Disease Information from the web (PADI-web)
3.2.3 Retrospective study of the novel coronavirus disease (COVID-19) in China using PADI-web
3.3. Development of an online data collection tool for mass gathering surveillance
3.3.1 Context of mass gatherings
3.3.2 Disease surveillance in the Pacific
3.3.3 Electronic disease surveillance
3.3.4 SAGES tool
3.3.5 Application of SAGES to enhance disease surveillance during the Micronesian Games in the Federated States of Micronesia in 2014
3.4. Section discussion
A pluridisciplinary approach to data analysis in the context of risk characterization
4.1. Introduction
4.2. The analysis of the circulation of arboviruses in the Pacific (2012-2014)
4.2.1 Introduction to arboviruses in the Pacific
4.2.2 Context of the study
4.2.3 Material and methods
4.2.4 Main results
4.2.5 Study discussion
4.3. The estimation of the spread rate of lumpy skin disease in the Balkans (2015-2016)
4.3.1 Lumpy skin disease
4.3.2 Modelling the spread rate of infectious diseases
4.3.3 Context of the study
4.3.4 Material and methods
4.3.5 Main results
4.3.6 Study discussion
4.4. Section discussion
Discussion
5.1. Summary of my contributions
5.2. Defining epidemic intelligence
5.3. Identification of drivers
5.4. Integrating different disciplines and expertise
5.5. Other levels of integration
5.6. Strategies to facilitate integration
Perspectives
6.1. Perspectives relating to the case studies
6.2. Exploring new data streams
6.3. Integrating new technologies
6.4. Disease x: anticipating the emergence of unknown viruses
6.5. From reactive to proactive
References