Get Complete Project Material File(s) Now! »
Chapter 3 Initial work on the thermal melt assay
This chapter will examine the initial work performed using the thermal melt assay. This was the first biological assay to be performed using the generation I dendrimers which were synthesised in chapter 2. The generation I dendrimers used in experiments mentioned in this chapter are shown in Figure 3.1.
Introduction to differential scanning fluorimetry
The cell binding assay described in Chapter 4 has some inherent difficulties. These cell assays re-quire healthy donated blood, from which the peripheral blood mononucleated cells (PBMC) need to be separated. PBMC are cultured for four days to develop into monocyte dendritic cells (MoDC) which express macrophage galactose binding lectin (MGL). MoDC have a limited lifetime and do not produce accurate results after being frozen and thawed. A fresh batch of PBMC needs to be cultured for every experiment, making it a costly and time consuming procedure. An alternative method for studying MGL binding during the optimisation of dendrimer design would be preferable.
An alternative to culturing PBMC to generate MGL expressing MoDC, would be to clone the DNA sequence for MGL and then to express the MGL protein in bacteria such as E. coli. This re-moves the need for human blood, and the MGL expressing bacteria can be generated from a stock whenever needed. Since the MGL from protein expression behaves the same before and after be-ing frozen and thawed, a stockpile of MGL truncations could be created and then defrosted when needed, as opposed to waiting for four days to perform a cell binding experiment.
MGL has previously been cloned and expressed independently by Suzuki et al. and Valladeau et al.[12] In 1996 Suzuki et al. first cloned the cDNA from a library derived from 1l–2 treated peripheral blood monocytes and then expressed the MGL recombinantly in COS-1 cells.[132] From the lysate of the cells, Suzuki et al. were able to separate out a protein of 38 kDa which was MGL, and then they went on to show that it bound to the TN antigen. In 2001, Valladeau et al. cloned a MGL splice variant which they named DC-asialoglycoprotein receptor.[133] The protein differs from MGL due to an insertion of 27 amino acids and a deletion of 3 amino acids in the neck domain.
In 2010 Drickamer expressed truncated MGL using an analogous T7 promoter system. With this system it was demonstrated that the presence of extended binding sites within a single carbohydrate recognition domain (CRD) can enhance binding interaction with branched glycans.[134] Drickamer went on to show that the presentation of branched glycans which contain GalNAc on a glycopro-tein surface increases binding affinity by 15–to 20–fold, possibly due to low-specificity interactions with the surface of the protein or restriction in the conformation of the glycans. Recently in 2013 Drickamer et al. cloned and expressed portions of the α–helical neck region and the globular CRD regions of MGL.[14] These fragments were used to confirm that MGL was a trimeric cluster due to its similarities in sequence with serum mannose-binding protein.
Truncations of MGL can be expressed by bacteria. It was hoped that truncations of MGL could be used in a thermal melt assay as a pre–screening tool to select ligands before submitting them for the cell binding assay. When this project started it was unknown whether this approach would be an accurate tool for predicting dendrimer binding to MGL, as it was unknown whether the MGL truncations would behave in a similar fashion to the presumably trimeric MGL found on MoDCs.
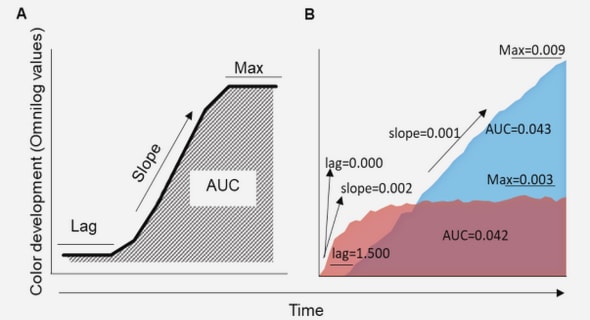
The thermal melt assay uses Differential Scanning Fluorimetry (DSF) to show the strength of binding of the compounds to MGL in a semi–quantitative manner. The better a ligand binds to the protein, the more stable the protein becomes, and therefore more thermal energy is needed to denature the protein. A compound which binds strongly will stabilise the protein and therefore will need a higher temperature to break up the tertiary structure compared to the protein on its own. The DSF experiment requires a fluorescent dye (SYPRO orange dye) which binds to hydrophobic amino acids of denatured proteins. As the temperature rises, the protein unfolds and hydrophobic amino acids which normally reside in the protein’s interior are exposed. The dye binds to these amino acids, and this results in a fluorescent signal. The DSF experiment produces plots in which the fluorescence intensity of the SYPRO Orange dye interacting with the peptide is plotted against temperature. The point of inflection on this graph indicates the temperature at which the protein denatures or « melts » (Figure 3.2).
As the point of inflection is often hard to see on the plots, the derivative curve (rate of change of fluorescence intensity against temperature) is normally used, since the point of inflection has become a minimum (Figure 3.2). The appendix of this thesis includes all of the DSF curves and derivative plots that were measured for this project. The melting points were obtained from the minima of the derivative graphs. The bar charts in this chapter were derived from the derivative graphs found in the appendix.
MGL is known to bind to GalNAc,[29, 30, 31] therefore GalNAc was used as a positive control for the thermal melt assay while the protein–only samples were used as a negative control. A compound will be considered a strong binder if the melting point is greater than that with GalNAc. Compounds which induce a lesser positive shift in the melting point than that of the GalNAc–MGL binding pair will be considered weaker binders. Compounds with zero or negative shift in melting point will be considered not to bind in a positive fashion (they may be destabilising the MGL or not interacting at all).
When preparing to run the thermal melt assay, the MGL sample was kept on ice to reduce the chances of the protein denaturing or aggregating before the experiment. The stock solution of pro-tein in buffer was made and then pipetted between each sample to maintain a consistent concentra-tion of MGL. To each sample of MGL, the dendrimers were added and then the solution was diluted to the desired concentration with buffer solution. Five minutes before the experiment was run the fluorescent dye was added to each sample to reduce the chance of non-specific dye binding to the protein before the DSF. A sample of each additive (the dendrimers/peptides/GalNAc monomer) was run with only the buffer, as well as with the buffer and the dye, to monitor for unwanted fluorescent signalling which may bias the test. Fortunately none of these compounds interacted with the dye or the buffer to produce a fluorescent signal.
The four MGL proteins
To perform the thermal melt assay, the MGL truncations were expressed recombinantly. Figure 3.3
[135] shows the complete MGL amino acid sequence and the four MGL truncation sequences which were expressed for this project. The MGL truncation sequences are: MGL 144 (amino acid 144 to 316, 21 kDa), MGL 155 (155 to 316, 20 kDa), MGL 166 (166 to 316, 19 kDa), and MGL 176 (176 to 316, 18 kDa). We chose to express the truncated sequences in bacteria rather than produce them by chemical synthesis for the following reasons:
• The long protein sequence would require substantial time to optimise the synthesis and fold-ing of MGL if it was made using solid phase peptide synthesis.
• Expression using bacteria is cheaper and faster than solid phase synthesis, and also produces proteins in larger quantities.
• The MGL truncations have been expressed before by researchers in the Squire group at the University of Auckland. The extraction and the folding have already been optimised to give correctly folded protein.
The biggest MGL truncation is MGL 144 which includes the largest portion of the coiled coil domain. The length of the coiled coil domain decreases with each subsequent truncation to the point that MGL 176 does not include any of the coiled coil domain. We postulate that the longer the coiled coil domain in the truncation, the more likely the MGL truncation forms trimeric MGL sequences that we hypothesise to be the most biologically relevant. Using this logic, we expect MGL 176 to be a monomer, while the larger sequences such as MGL 144 and MGL 155 is expected to form trimers.
Expression of the four MGL constructs
The four MGL truncation domain constructs were provided by Dr. Christopher Squire (SBS, UoA). These four sequences had already been inserted into the pDEST17 Gateway vector (Invitrogen) and transformed into E. coli BL21(DE3). The transformed bacteria were grown on agar plates supple-mented with 56 nM ampicillin at 37 ◦C and left to incubate for 18 hours. pDEST17 confers ampi-cillin resistance and has the protein encoding sequence under a T7 promoter. This should reduce the amount of E. coli produced which do not express the MGL sequences. A single colony was taken from the agar plate after the incubation and was then transferred into liquid media to continue to be expressed.
Initially we used the isopropyl β–d–1–thiogalactopyranoside (IPTG) to trigger the transcription of the lac operon and induce protein expression. IPTG induction was very low yielding, and there-fore we switched to using Terrific Broth un-induced expression which expressed more protein. After expression, the supernatant was separated from the E. coli solids by centrifugation, and the resulting pellet was re-suspended in a tris buffer solution. The suspension was put through a constant cell disruption system instrument to lyse the cells.
After the cell lysis, the mixture underwent centrifugation and the supernatant was separated from the pellet. The sodium dodecyl sulfate (SDS) polyacrylamide gel electrophoresis (PAGE) of the four MGL sequences (Figure 9.22, appendix), showed that the MGL is located in the pellet rather than the supernatent as expected from previous expression experiments. The pellet was re– suspended in a buffer containing guanidinium, calcium chloride and β–mercaptoethanol (β–ME) to solubilise the inclusion bodies which contain the MGL. The solution was agitated overnight at 4 ◦C. The solubilised MGL was then separated from the rest of the cell debris by centrifugation. Unfortu-nately SDS–PAGE analysis can not be performed at this stage, as the guanidinium disrupts the shape of the gel. The concentration of the MGL in solution was normally measured using a Nano–drop fluorospectrometer. Unfortunately again due to the guandinium, the Nanodrop fluorospectrometer does not give accurate results for the concentration measurement.
The MGL at this stage was in an unfolded state, and the protein must be allowed to refold into its functional 3D shape. The MGL solution was added very slowly to a large excess of refolding buffer at 4 ◦C and left to stir overnight. The rapid dilution of protein should dilute the β–ME and guanidinium enough to allow the protein to refold to its native conformation. A fraction of the protein deposits out overnight and the solution goes slightly cloudy.
The refolding buffer with MGL was then filtered and concentrated from 500 mL down to 200 µL. The filter allows proteins smaller than 10 kDa to pass through but stops the larger proteins (which include each of our MGL truncations) from passing through. The more concentrated the solution, the more protein precipitates. When the solution was 200 µL in volume, a Nanodrop fluorospec-trometer measurement was performed, which revealed the concentration of the MGL. The super-natant before size exclusion chromatography produced an SDS–PAGE trace which was a lot cleaner than the initial expression. An example of SDS–PAGE for MGL 144 truncation before size exclusion chromatography is included in Figure 3.4.
Size exclusion chromatography
The concentrated supernatant contains a mixture of proteins which remain after the rapid dilu-tion and filtering. To separate the desired MGL truncation, size exclusion chromatography was performed. Initially we tried to purify the samples on a Superdex 75 10/300 column with just the refolding buffer. Unfortunately the column blocked repeatedly due to the MGL truncations precipi-tating. This problem could be solved by adding a small amount of BME (β-mercaptoethanol) to the refolding buffer. The profile of the MGL traces is shown in the appendix (Figure 9.24 to Figure 9.26).
While all of the smaller truncations gave a similar trace, MGL 144 eluted in the void volume when we attempted to purify using the Superdex 75 10/300 column (Figure 3.4). To solve this issue, a Superdex 200 10/300 column was used instead (Figure 3.5). The fractions for each run were collected and then tested using SDS–PAGE to identify the fractions with the desired protein. These fractions were concentrated to 200 µL and then their protein concentration was measured using a nanodrop fluorospectrometer.
The protein was then diluted with 20% glycerol and flash–frozen in liquid nitrogen at -80 ◦C to prevent degradation. The expression of the MGL truncations was good but the final yields were always low because of the insolubility of the protein during the purification process and especially during the refolding steps. Due to time limitations, we did not explore the use of other refolding buffers, the buffer used was known to work from previous unpublished work from the Squire group. If the yield of the expression from MGL truncations were to be improved, the refining of the refold-ing conditions would be the primary focus.
The Superdex 75 10/300 column is suitable for small proteins, and it is suitable to resolve monomeric MGL truncations which have a molecular weight of around 20 kDa. MGL 144 was the last truncation to be purified using size exclusion chromatography. All of the smaller truncations were purified without issue. Therefore the size exclusion chromatography trace for MGL 144 shown in Figure 3.4 with only one main peak was unexpected.
A calibration experiment was performed using proteins of known mass and the calibration pro-tein of 60 kDa eluted near the void volume, leading us to believe that MGL 144 (monomer of 21 kDa) may form trimeric structures (due to containing the longest portion of the coiled coil do-main). The SDS-PAGE shown in Figure 3.4 showed protein with masses between 50 and 70 kDa present, which could correspond to the molecular weight of the 63 kDa MGL 144 trimer. The MGL 144 monomer was also separated during the purification, which suggests that the formation of the quaternary structure did not go to completion with the current refolding conditions producing a monomer/trimer equilibrium.
When MGL 144 was purified using the Superdex 200 10/300 column, a better resolved size exclusion trace was generated (Figure 3.5) when compared to the Superdex 75 10/300 column (Fig-ure 3.4). The MGL 144 trace has a similar profile to the other smaller MGL truncations which were purified on the Superdex 75 10/300 column.
Size exclusion chromatography multi–angle laser light scattering (SEC-MALS) was briefly con-sidered as a way to prove the trimeric structure of the MGL 144 protein. Unfortunately, the refolding buffer which contains a high concentration of glucose made calibration impossible. Due to time con-straints, alternative solvent systems weren’t explored, and therefore SEC-MALS wasn’t performed. Future research could be directed towards the optimisation of solvent systems for: MGL refolding, purification, SEC-MALS and crystallisation as this could reveal the precise quaternary structure.
The initial thermal melt assay
Dr Julie McIntosh provided the initial samples of MGL 176 protein which were used for the first DSF experiments. MGL 176 is expected to be monomeric as it contains the carbohydrate binding domain of MGL but does not contain the coiled coil domain which induces oligomerisation. The coiled coil domain is thought to be important for the formation of trimeric clusters in MGL. We hoped to use MGL 176 as a model system to compare to the full MGL sequence, as it contained the full carbohydrate binding domain which was where we were expecting the GalNAc dendrimers to bind.
The initial thermal melt assay was a titration experiment using monomeric GalNAc and the gen-eration I dendrimers: 21, 2, 23 and 25. The results are shown in Table 3.1 and Figure 3.6. MGL is known to bind to GalNAc, and was therefore used as a positive control for the experiment.[29, 30, 31] The GalNAc induced an increase in the melting point of MGL (from 2 ◦C at the lowest concen-tration to 4 ◦C at the highest concentration). The dendrimers 21, 2 and 23 all performed better than GalNAc. The shifts of over +5 ◦C by the three dendrimers which bound to MGL (compounds 21, 2 and 23) were very encouraging. Dendrimer 2 performed particularly well, and was able to induce an 8.5 ◦C shift in melting point at a 300 µM concentration. Dendrimer 25 performed poorly, and induced either no shift or a negative shift in the melting point.
Thermal melt assay titration
The original DSF experiments followed a procedure developed in the laboratory of Dr Christopher Squire (SBS, UoA); unfortunately this showed a limited range of concentration for additives to in-teract with MGL. The aim of this work was to improve the experiment to show how the difference in concentration affects the binding to MGL and hence find the optimum concentration of the den-drimers for the thermal melt assay experiments. The concentration range used for the titration DSF was from 1 nM of the additive, which was also the lowest concentration used for additives in the cell binding assay, to 1 mM the maximum concentration that could be used in multiple experiments, without rapidly exhausting all of the dendrimer stocks. GalNAc was once again used as the positive control.[29, 30, 31] Two samples per additive were used in the DSF procedure originally developed by Squire et al. This was increased to three samples per additive in order to increase the statistical relevance.
The average melting points for the titration experiment are shown in Table 3.2 and Figure 3.7. The results indicate that when GalNAc binds to MGL in the concentration range of 1 nM to 100 µM, the melting point increases by roughly 2 ◦C. Incubations at 1 mM of GalNAc result in the melting point of MGL 176 increasing significantly by 6 ◦C. Dendrimer 21 always had at least a 1 ◦C larger shift in melting point increase than GalNAc over all concentrations. Dendrimer 21 also showed that the critical concentration for larger shifts in melting point occur was at 1 mM, where it induced much larger shifts in melting point (10 ◦C) than GalNAc (6 ◦C). The error bars for the experiment are small as there is a small spread in the data which resulted in the small standard deviation. Since the largest shifts in melting point for the titration experiment consistently occurring at 1 mM it was decided that the subsequent experiments for the thermal melt assay would use this concentration when comparing different additives.
The curve of the titration shown in Figure 3.7 shows that the binding to MGL 176 has not become saturated yet. Saturation would normally be indicated by a plateau forming in the plot, at which increasing the concentration of the additive will not increase in the melting point temperature. For the practical reason of dendrimer conservation, the experiment did not use larger concentrations, as this would deplete the stock too quickly.
The initial thermal melt assays showed that the GalNAc–containing generation I dendrimers would interact in a positive fashion with the carbohydrate recognition domain of monomeric MGL, and that the strength of binding was in proportion to the concentration of the dendrimer incubated with MGL. The next step would be to test the binding of the dendrimers to cells which express MGL.
Contents
0.1 Abstract
0.2 Preface
0.3 Acknowledgements
0.4 Abbreviations
1 Introduction
1.1 Targeting dendritic cells
1.2 C–type lectins
1.3 MGL
1.4 The role of MGL in diseases
1.5 Peptide vaccines and drug delivery
1.6 Click chemistry
1.7 Examples of TN antigen vaccines
1.8 Aim
2 Initial synthesis of generation I and II dendrimers for targeting MGL
2.1 The design of a dendrimer
2.2 Retrosynthesis of the dendrimer-glycan
2.3 Synthesis of building blocks: Fmoc–azidoalanine and α–propargyl GalNAc(Ac)3
2.4 Synthesis of the generation I dendrimers
2.5 The CuAAC reaction
2.6 The CuAAC reaction with non–protected α–propargyl GalNAc
2.7 Generation II dendrimers
3 Initial work on the thermal melt assay
3.1 Introduction to differential scanning fluorimetry
3.2 The initial thermal melt assay
3.3 Thermal melt assay titration
4 Ability of GalNAc compounds to bind MGL–expressing human cells at 0 ◦C
4.1 Introduction
4.2 Gating strategy for flow cytometry experiments.
4.3 MGL–expression and selection of the MGL negative LG2 cells for the cell binding assay109
4.4 Optimisation at 0 ◦C
4.5 Optimised 0 ◦C conditions
5 Synthesis of generation III, IV and V MGL targeting compounds
5.1 The importance of the X, Y and Z chain to binding.
6 Thermal melt studies on the four MGL truncations
6.1 Thermal melt assay for the four truncations of MGL using the library of dendrimers
7 The MGL–expressing MoDC binding assay at 37 ◦C
7.1 MUC1 sequences synthesised by Dr Lee
7.2 Optimisation at 37 ◦C
7.3 Results from optimised conditions
7.4 Discussion
7.5 Conclusion
7.6 Does the thermal melt assay predict the results of the cell binding assay?
7.7 Targeting MGL–expressing cells in peripheral blood mononucleated cells
7.8 Results of binding of dendrimers to white blood cells in human blood
7.9 Discussion of binding of dendrimers to white blood cells in human blood
7.10 Summary of the thesis
7.11 Future work
8 Experimental
8.1 Experimental conditions and analytical parameters
8.2 Synthesis of unnatural amino acids and carbohydrate building blocks
8.3 General methods for the synthesis of dendrimer and peptide scaffolds
8.4 General amino acid coupling methods
8.5 Cleavage methods
8.6 Synthesis of dendrimer and peptide ligands
8.7 Cell binding assay
8.8 Thermal melt assay
9 Appendix
9.1 Chemistry appendix
9.2 Thermal melt assay appendix
9.3 Size exclusion chromatography traces
9.4 Thermal melt
9.5 Cell binding assay appendix
Bibliography
GET THE COMPLETE PROJECT