Get Complete Project Material File(s) Now! »
Immunohistochemistry on thymic sections
Frozen thymic sections (7μm) were fixed in ice-cold acetone for 20 minutes. Immunofluorescent staining was performed by one hour incubation at room temperature with antibodies listed in table S1b. Images were acquired with a Zeiss Axio Observer Z1 Inverted Microscope using 40X and 63X magnifications.
Flow Cytometry
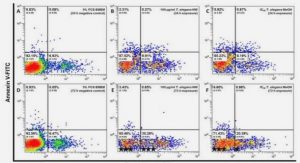
Cell populations were identified according to their characteristic FSC/SSC profile and by staining with the following antibodies: B cells with an anti-CD19 antibody, T-cell subsets with anti-CD4 or anti-CD8 antibodies and monocytes with an anti-CD14 antibody. Myeloid DCs (mDCs) were defined as CD14-CD11c+ cells and plasmacytoid DCs (pDCs) as CD14- CD123+ cells. For the analysis of intracellular CXCR4, CXCR7 and SDF-1, PBMCs were stained for surface CD11c and CD14 antigens to identify mDCs, fixed and permeabilized with an eBioscience permealization Kit (Montrouge, France) and labeled with anti-CXCR4, anti- CXCR7 or anti-SDF-1 antibodies. All antibodies are listed in table S2. Staining conditions for all antibodies were 30 min at 4°C in the dark. Cells were then fixed and acquired using BD FACScalibur. Data was analyzed with FlowJo analysis software (Treestar, San Carlos, CA). For cell sorting, PBMCs were stained with anti-CD14, anti- CD11c, and anti-CD3 antibodies to identify mDCs, monocytes and T cells, and run on a 4- laser FACS Aria high-speed cell sorter using FACS Diva acquisition software (both from BD Biosciences).
Laser-capture microdissection
Microdissection was performed as previously described (19). To label HEVs, sections were consecutively stained with anti-PNAd carbohydrate epitopes, biotinylated mouse anti-rat and streptavidin-horseradish peroxidase. Antigen was visualized with the DAB+ system (K3467, DAKO) and sections were counterstained with hematoxylin. HEV and medullary zones were separately microdissected before RNA extraction.
Reverse transcription and real-time PCR
Total RNA was extracted as previously described (20)! »#$% »&' »()* »+,-« ./0/.-/ »1.,2-3.45/6″ for 1 hour at 42°C using AMV (Eurobio, Courtaboeuf, France) with oligo-dT (Invitrogen). RNA from laser-capture microdissection was extracted with RNeasy Micro Kit (74004, Qiagen, Courtaboeuf) and reverse-transcribed using SuperScript II with oligo-dT and random primers (Invitrogen). Real-time PCR reactions were performed on the LightCycler apparatus as previously described (20). The primer sequences (Eurogentec, Angers, France) were as follows: (1) for SDF-#7″ 89:;<- CACTGTGGCTAACATTG-3’, R:5’- AGGTCCTGGTGGTATT-3’), (2) for SDF-#= » 89:;<-TAGTCAAGTGCGTCCACGAG-3’, R:5’- ACACACAGCCAGTCAACGAG-3’), (3) for SDF-#7>= » 89:;<- CTTTAGCTTCGGGTCAATGC-3’, R:5’-CAGCCTGAGCTACAGATGC-3’), (4) for 28S (F5’-CGGGTAAACGGCGGGAGTAA-3’, R:5’-GGTAGGGACAGTGGGAATCT-3’), (5) for GAPDH (F5’-GATGATGTTCTGGAGAGCCC-3’.
Analysis of chemokine expression on thymic HEVs in MG
In SLOs and inflamed tissues, HEVs display chemokines on their surface which mediate adherence and transmigration of peripheral cells to the target tissue (9). We therefore analyzed the chemokine expression pattern on thymic HEVs in MG. Sections of hyperplastic MG thymus were co-labeled with an anti-PNAd antibody for HEVs and antibodies against selected chemokines: CXCL9, CXCL10, CXCL11, SDF-1, CXCL13, CCL19, CCL21 and RANTES. The analyzed chemokines were chosen by two criteria: a reported expression on HEVs in SLOs or other inflamed tissues, and a reported dysregulation in MG thymus, as detailed in table S2. From all these chemokines, we exclusively observed SDF-#7>= » expression on the lumen side of HEVs (Fig. 3A) but not SDF-#? »86,1, »2&1″-@&+2A! In non-MG adults with a few HEVs, we observed that HEVs also displayed SDF-#7>=! »Analyzing mRNA expression of SDF-! »#and -$, the two main SDF-1 isoforms expressed in the thymus (24), we did not observe significant differences in thymic extracts (Fig. S1A).
Peripheral cells around HEVs are partially CXCR4+
As SDF-1 mediates cell recruitment by binding to its receptor CXCR4, we analyzed thymic CXCR4 expression in MG. Globally, CXCR4 mRNA expression levels did not differ between the thymus of MG patients and non-MG adults (Fig. S1B). The fact that the global CXCR4 level did not vary could be due to the dominant CXCR4 expression by TECs and thymocytes, which may mask differences between MG an non-MG adults. However, when we performed co-staining for HEVs, peripheral cells and CXCR4, we observed that some of the B cells, monocytes/macrophages, mDCs and pDCs that localized around HEVs, expressed CXCR4 suggesting that they could have been recruited through binding to SDF-1 on HEVs. We were also able to detect CXCR4+ monocytes/macrophages (Fig. 4B, lower row) and pDC (Fig. 4B, lower row) inside HEVs strongly supporting this hypothesis. CXCR7 is a second receptor for SDF-1, whose role in cell migration processes is debated (13, 14). Analyzing CXCR7 mRNA levels by real-time PCR, we did not observe differences between MG and non-MG thymus. By immunohistochemistry on thymic sections from MG, we only detected CXCR7 on GClike structures (data not shown).
Table of contents :
I. STATE OF THE ART
1. MYASTHENIA GRAVIS
1.1 GENERALITIES
1.1.1 HISTORY
1.1.2 EPIDEMOLOGY
1.1.3 SYMPTOMS AND DIAGNOSIS
1.2 PATHOPHYSIOLOGICAL MECHANISMS IN MG
1.2.1 THE NEUROMUSCULAR JUNCTION UNDER PHYSIOLOGICAL CONDITIONS
1.2.2 PATHOGENIC MECHANISMS IN MG
1.2.3 GENETIC PREDISPOSITION
1.2.4 TREATMENTS
1.3 EXPERIMENTAL AUTOIMMUNE MG
1.4 NEW POTENTIAL THERAPIES
2. THYMUS
2.1 STRUCTURE AND FUNCTION
2.1.1 GENERALITIES
2.1.2 THYMIC CELLS
2.1.3 T CELLS: MATURATION BY NEGATIVE AND POSITIVE SELECTION
2.2 PATHOLOGICAL ROLE
2.2.1 THYMIC DISORDERS
2.2.2 THYMOMA-ASSOCIATED MG
2.2.3 THE HYPERPLASTIC THYMUS IN MG
3. CHEMOKINES
3.1 GENERAL PROPERTIES
3.1.1 STRUCUTRE AND CLASSIFICATION
3.1.2 CHEMOKINE RECEPTOR SIGNALING
3.2 PHYSIOLOGICAL ROLE OF CHEMOKINES
3.2.1 LYMPHOCYTE HOMING
3.2.2 THYMOPOIESIS
3.3 PATHOLOGICAL ROLE OF CHEMOKINES
3.3.2 CHEMOKINES IN MG THYMUS
II. AIM OF THE STUDY
III. RESULTS
Article 1
Article 2
Article 3
Article 4
IV. GENERAL CONCLUSION
V. PERSPECTIVES
VI. REFERENCES